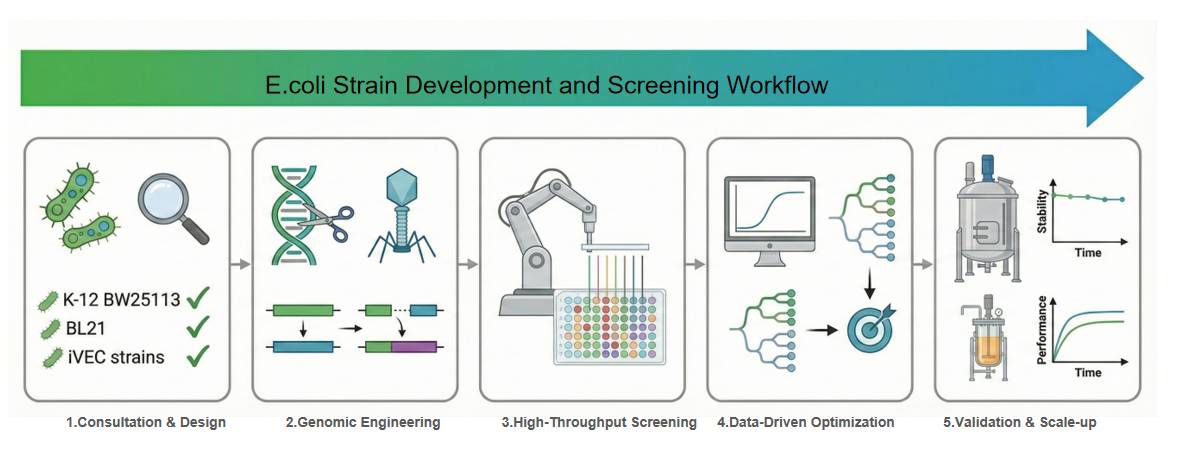
Application Study 1: High-Throughput Screening (HTS) Platforms and Industrial Applications
This report details how CDMOs (Contract Development and Manufacturing Organizations) such as KBI Biopharma utilize high-throughput screening (HTS) and automated micro-fermentation platforms to accelerate strain development. By using 96-well plates or droplet technology, researchers can rapidly evaluate strain growth rates and product generation capabilities under small-volume conditions. The automation system integrates liquid-handling robots, significantly reducing the variability and time costs associated with manual operations. This is particularly critical when screening for recombinant proteins that are difficult to fold or require disulfide bonds, such as antigen-binding fragments (Fabs), cytokines, and enzymes.
(Reference: Brian Gazaille with Erik Nordwald, 2023)
Application Study 2: Data-Driven Strain Design and Adaptive Evolution
This research demonstrates how to utilize Adaptive Laboratory Evolution (ALE) databases for computer-aided strain design. By aggregating large amounts of mutant data, researchers can identify gene mutations that perform exceptionally well in specific environments (such as high salt or acidic pH). Furthermore, metabolic network models are used to predict which gene knockouts or overexpressions can improve the flux of target metabolic pathways. This allows for more targeted gene editing—rather than blind random mutagenesis—and is applicable to both biofuels and high-value pharmaceutical intermediates.
(Reference: Phaneuf, Patrick V. et al., 2021)
Application Study 3: Strain Library Resources and Functional Genomics Toolkits
This report introduces the Keio Collection and iVEC strain libraries, which are fundamental tools for genome-wide screening and functional analysis. The Keio Collection contains single-gene knockout mutants for 3,985 non-essential genes, providing a unified background strain (E. coli K-12 BW25113) for systematic analysis. Additionally, iVEC strains support seamless cloning, facilitating the rapid construction of recombinant plasmids. Using a unified strain background significantly reduces experimental variability and improves the comparability of screening results, allowing for the rapid identification of key genes related to target product synthesis.
(Reference: MEXT National BioResource Project, 2022)
Application Study 4: Targeted Gene Knockout and Metabolic Pathway Reconstruction
This study demonstrates the creation of a metabolic "blank" platform for the biosynthesis of non-natural amino acids through specific gene knockouts (such as ΔtrpR, ΔpheA, and ΔtyrA). By knocking out the host's natural metabolic pathways to prevent substrate competition, heterologous pathways expressed by exogenous genes can monopolize resources for target product synthesis. This strategy successfully incorporated halogenated tryptophan derivatives, showcasing the platform's potential in synthesizing complex natural product precursors and its ability to adapt to diverse metabolic engineering needs through simultaneous multi-gene knockouts.
(Reference: Zhang et al., 2021)