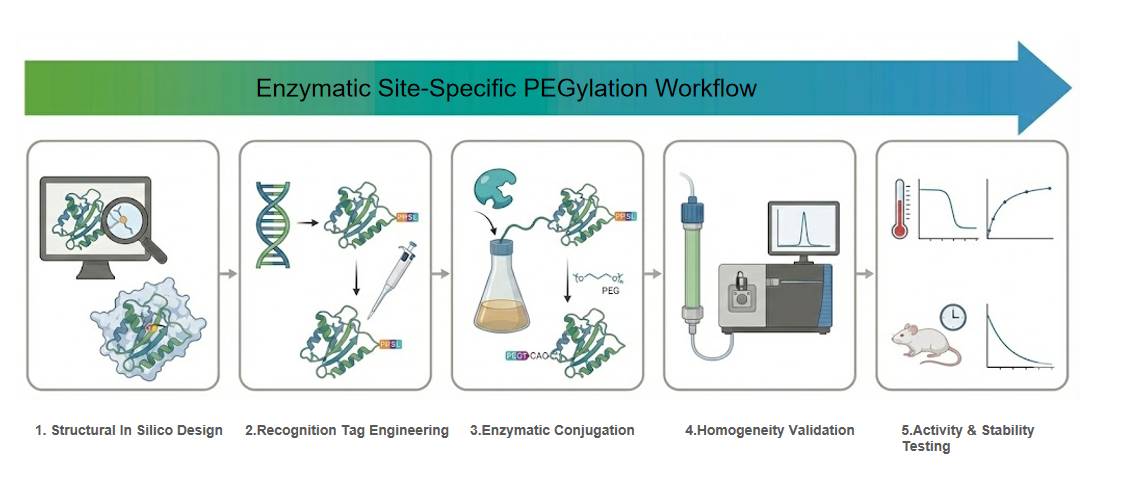
Application Study 1: High-Efficiency Plastic Degradation (PETase)
Industrial PET hydrolases often suffer from poor stability. By utilizing computer-aided enzymatic PEGylation, technical benchmarks increased PETase thermal stability by 3.88°C and boosted degradation rates by 3.5-fold at 30°C. This provides a scalable solution for biological plastic waste processing.
(Reference: ACS Applied Materials & Interfaces, 2024)
Application Study 2: Chemoenzymatic Glyco-Tagging for "Biobetters"
Native proteins can be upgraded using glycosyltransferases to attach precise glycan/PEG tags. This strategy significantly enhances PK/PD properties while lowering immunogenicity compared to random chemical methods, allowing for the rapid development of superior biosimilars.
(Reference: Nature Biomedical Engineering, 2024)
Application Study 3: Sustained Immunosuppression via IL-2 Modification
Interleukin-2 is critical for treating autoimmune diseases. Enzymatic site-specific PEGylation of IL-2 resulted in sustained activation of regulatory T cells (Tregs) and vastly improved in vivo stability, offering new potential for transplant rejection therapies.
(Reference: Nature Biomedical Engineering, 2021)
Application Study 4: Improving Therapeutic Efficiency of L-asparaginase
For anti-cancer enzyme treatments, subunit stability is paramount. Enzymatic PEG-crosslinking creates a uniform, stable conjugate that bypasses hemodynamic barriers more effectively, extending circulatory time while reducing side effects.
(Reference: bioRxiv / Enzyme Subunit Crosslinking, 2018)