April 25th is DNA day, a big day in the history of biology, because 68 years ago, on April 25th, 1953, James Watson, Francis Crick, and their colleagues published a paper on the double helix structure of DNA in Nature. Since then, the mystery of DNA structure that has plagued generations has been solved, and the development of life sciences has entered the stage of molecular biology. The discovery of the double helix structure of DNA has greatly contributed to the flourishing of life sciences and also pointed out the initial direction for DNA synthesis. Today, we will discuss the evolutionary history of DNA synthesis technology and how it has benefited humanity.
What is the use of DNA synthesis? How to synthesize DNA?
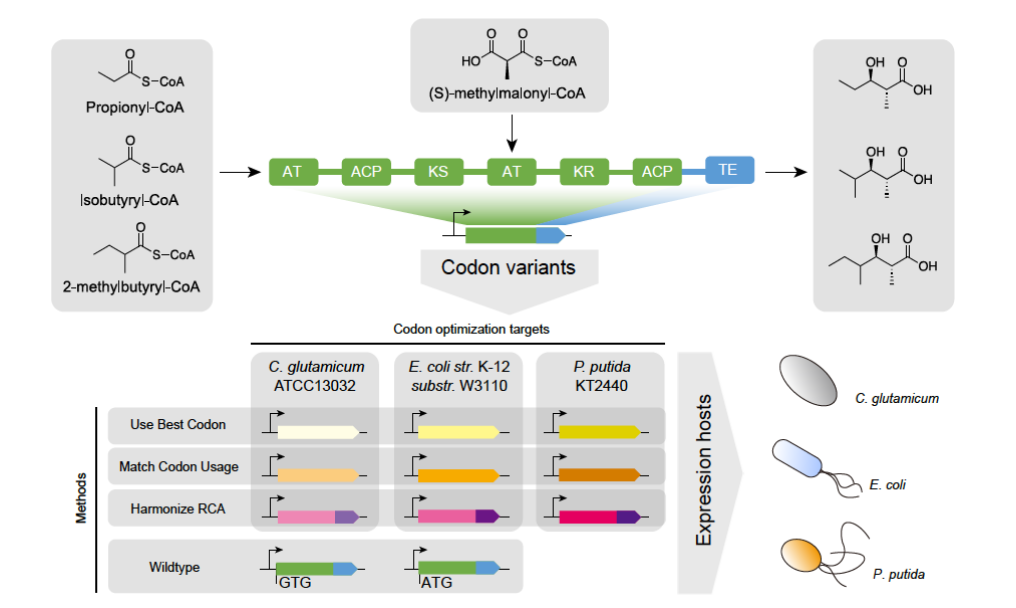
Technological progress is often application driven, and to understand the development history of DNA synthesis technology, we need to first understand the uses of DNA synthesis. In recent years, synthetic biology has rapidly emerged, in which codon optimization, metabolic engineering, gene assembly, and so on all rely on DNA synthesis. In addition, DNA synthesis also plays an important role in fields such as gene and cell therapy, molecular diagnosis, antibody and protein engineering, and will not be listed here.

We know that in living organisms, the way DNA replicates is through semi preservation replication, which means that when DNA replicates, two strands are released separately as templates. In the presence of primers, DNA polymerase, and dNTP, two new strands of DNA are synthesized according to the law of complementary base pairing. In newly synthesized DNA molecules, one strand is newly synthesized and the other strand is the original, hence it is called semi retained replication. In the laboratory, the synthesis of cDNA fragments through processes such as RNA extraction and reverse transcription is also a relatively traditional and classic DNA synthesis method
With the increasing demand for DNA synthesis, the advancement of DNA synthesis technology has been spurred, and direct synthesis of DNA has become a reality. The basic principle is to first combine long DNA single strands, which have complementary pairs, and then obtain complete sequences through PCR amplification .
The Historical Evolution of DNA Synthesis
Since the discovery of the double helix structure of DNA, DNA synthesis has gradually emerged on the historical stage. DNA synthesis has gone through a long and tortuous development, from the initial form of single stranded oligonucleotide chains to the current form of chromosome synthesis, making a leap forward. Overall, there are several landmark events in DNA synthesis:
Before 1970: Gene synthesis was only in the form of single stranded oligonucleotide chains. The synthesis of double stranded DNA (>100 bp) began to rapidly develop after 1970.
In 2002, the genome of poliovirus (approximately 7500 kb) was successfully synthesized, marking the beginning of the era of genome synthesis.
In 2010, Venter and Smith et al. transferred the artificially synthesized Mycoplasma mycoides genome into the host cells of Mycoplasma capricolum, marking a historic step in the creation of new cells through artificial gene synthesis.
In 2011, the yeast genome synthesis project (Sc2.0) began, and the synthesis of chromosomes 2, 3, 5, 6, 10, and 12 has been completed.
Future: The synthesis and assembly of DNA may involve the de novo establishment of complex microbial communities or cellular tissues.
picture
The evolutionary history of DNA synthesis technology
Behind the rapid progress in DNA synthesis and the achievement of iconic breakthroughs is the continuous innovation and iterative upgrading of DNA synthesis technology. There are many techniques for DNA synthesis, and the classification methods are also different, including those classified from a technical principle perspective, those classified according to a timeline, and so on. Generally speaking, mainstream DNA synthesis techniques can be roughly divided into four generations . The initial DNA synthesis was carried out using the phosphoramide triester synthesis method, which involves immobilizing DNA onto a solid-phase carrier to complete the synthesis of DNA strands . The second-generation DNA synthesis technology is based on chips, including inkjet, photochemical, and electrochemical methods. The third-generation DNA synthesis technology is an ultra-high throughput synthesis technique, which combines semiconductor and electrochemical methods. At present, the latest generation of DNA synthesis technology is enzymatic synthesis technology, mainly including microarray method, yeast in vivo DNA synthesis method, and linker mediated DNA synthesis method.
The technical process and related principles of DNA synthesis
The main process of gene synthesis includes the following four steps, with oligonucleotide synthesis and DNA assembly being the core;
DNA sequence design: By using computer software to design DNA sequences, these DNA sequences can form biological circuits or pathways.
Oligonucleotide synthesis: The DNA sequence is divided into smaller overlapping fragments (usually 200-1500bp), which are then further divided into a chemically synthesized set of single stranded oligonucleotides.
DNA assembly: Single stranded oligonucleotides are assembled into larger DNA fragments through various DNA assembly techniques and subjected to cloning validation.
Functional testing and feedback: The validated DNA sequence is transferred into cells for functional testing; Based on the results of functional testing, make changes to the design sequence for this round and repeat the testing cycle to form an iteration.

The synthesis of oligonucleotides: Currently, the main method used is the chemical synthesis of columnar phosphoramides; This process couples acid activated deoxyribonucleotide phosphoramide molecules onto a deoxyribonucleotide molecule fixed in a solid phase (controllable pore glass/CPP or polystyrene microbeads/PS beads). In the first synthesis cycle, the nucleotide chain extends from the first protected nucleotide molecule fixed on the surface.

These oligonucleotides can typically be synthesized up to 100nt. Longer nucleotide chains (up to 300-600 nt) can also be directly synthesized, but as the length increases, the yield of oligonucleotide synthesis will sharply decrease (random mutations are introduced during the synthesis process, and adenine is easily affected by purine removal, leading to main chain breakage).
DNA assembly: Generally, long stranded DNA is obtained by splicing and assembling pre synthesized oligonucleotides (usually 60-120 nt) as raw materials. Among them, ligase based assembly and polymerase chain reaction based assembly are the two most important methods.
Assembly method based on DNA ligase: Using DNA ligase to react complementary overlapping DNA strands to form longer fragments. The use of heat-resistant DNA ligases can prevent the formation of secondary structures in DNA and improve the success rate of assembly.

● DNA polymerase based assembly method: Polymerase chain reaction (PCR) is the most commonly used gene assembly method based on oligonucleotides. By using a one-step PCR assembly method, a set of oligonucleotide mixtures can be assembled into gene length DNA, but the product obtained by this method is usually a set of DNA with different lengths (Figure A). Another two-step assembly method based on PCR involves first breaking down the gene into a series of sub libraries, assembling them into fragments, mixing these fragments, and finally assembling them into complete genes (Figure B).
Microarray based DNA synthesis technology will become a future development trend
The biggest pain point of the gene synthesis industry at present is mainly the high cost, with a cost per base of 0.10-0.30 US dollars (the cost of 1kb genes is 100-300 US dollars), and there has been no significant decrease in the past 10 years. In addition, the automation level of DNA synthesis is not high, and multiple intermediate processes rely on manual operations. Therefore, reducing the consumption of synthetic reagents, increasing throughput, and improving the automation and accuracy of gene assembly processes are necessary to significantly reduce the overall cost of gene synthesis.
The microarray synthesis platform can significantly improve the synthesis flux of oligonucleotides, significantly reduce the amount of reagents required for synthesis, and thus greatly reduce costs. The cost of synthesizing oligonucleotides based on microarray platforms varies from $0.00001 to $0.0001 depending on the platform, oligonucleotide length, and synthesis scale. Compared to the cost of $0.05- $0.10 per base for columnar synthesis of oligonucleotides, chip synthesis of oligonucleotides has significant advantages in gene synthesis applications.

Nowadays, the moat of technology has formed, and only by mastering core technologies can one occupy a place in the DNA synthesis market. The DNA synthesis technologies mentioned above have their own limitations and technological barriers. CD Biosynsis has been deeply involved in the field of DNA synthesis for many years. Currently, CD Biosynsis has technology and patents in both DNA chip synthesis technology and ultra-high throughput synthesis, making it highly competitive in the industry.
We have reason to believe that with the advancement of DNA synthesis technology, the cost and threshold of DNA synthesis will be further reduced. While providing basic services for life science, it will also provide more assistance in revealing the mysteries of life. We celebrate DNA Day as a gratitude to the scientists who have made contributions to the discovery and application of DNA, as well as a hope for the future of life sciences to benefit humanity more.